PDMR-292921-168-R | Imaging characterization of a metastatic patient derived model of adenocarcinoma pancreas:
DOI: 10.7937/TCIA.2020.PCAK-8Z10 | Data Citation Required | 207 Views | Image Collection
| Location | Species | Subjects | Data Types | Cancer Types | Size | Status | Updated | |
|---|---|---|---|---|---|---|---|---|
| Abdomen | Mouse | 20 | MR, SR, Protocol | Pancreatic Adenocarcinoma | Clinical | Public, Complete | 2020/03/23 |
Summary
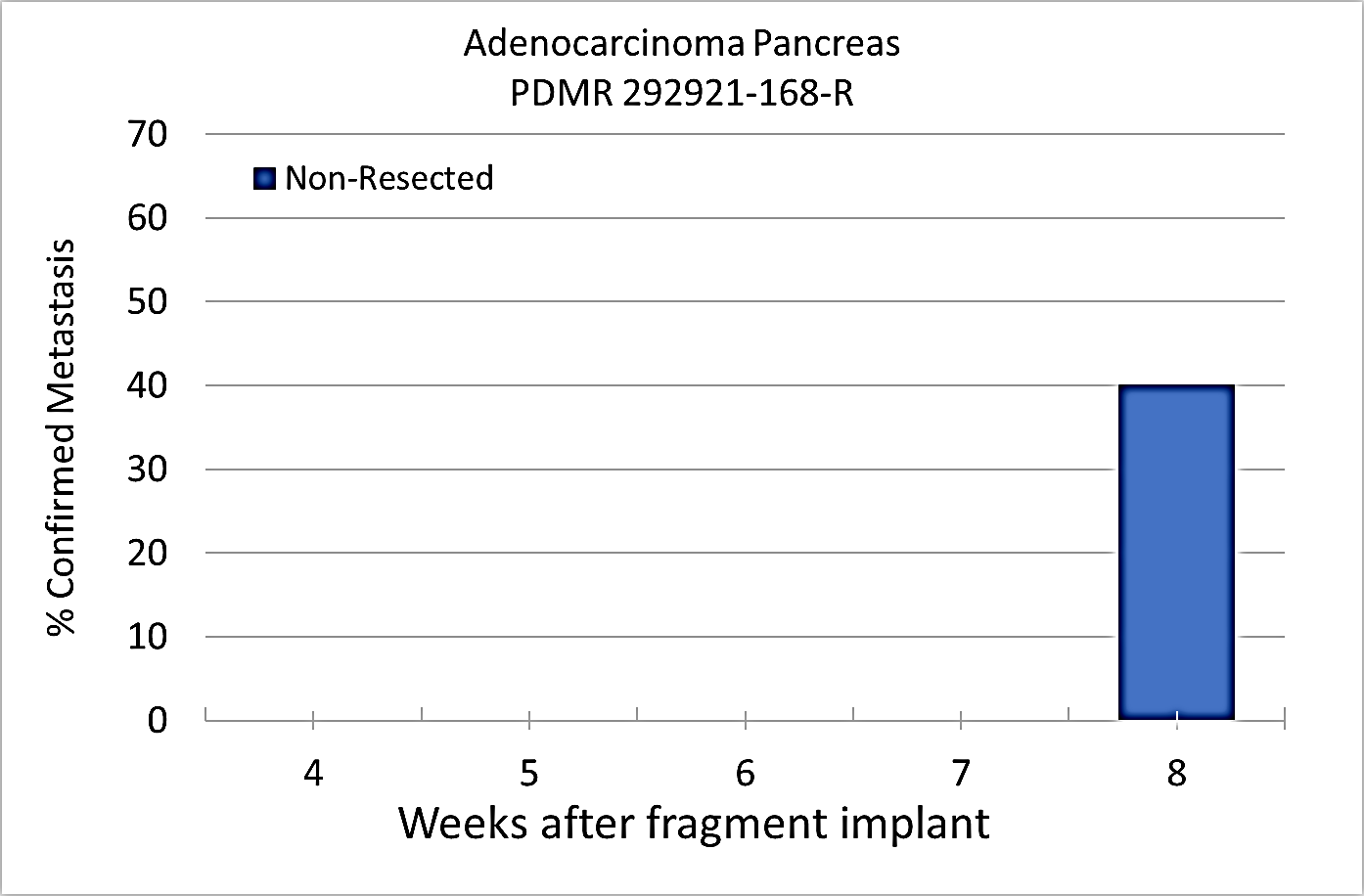
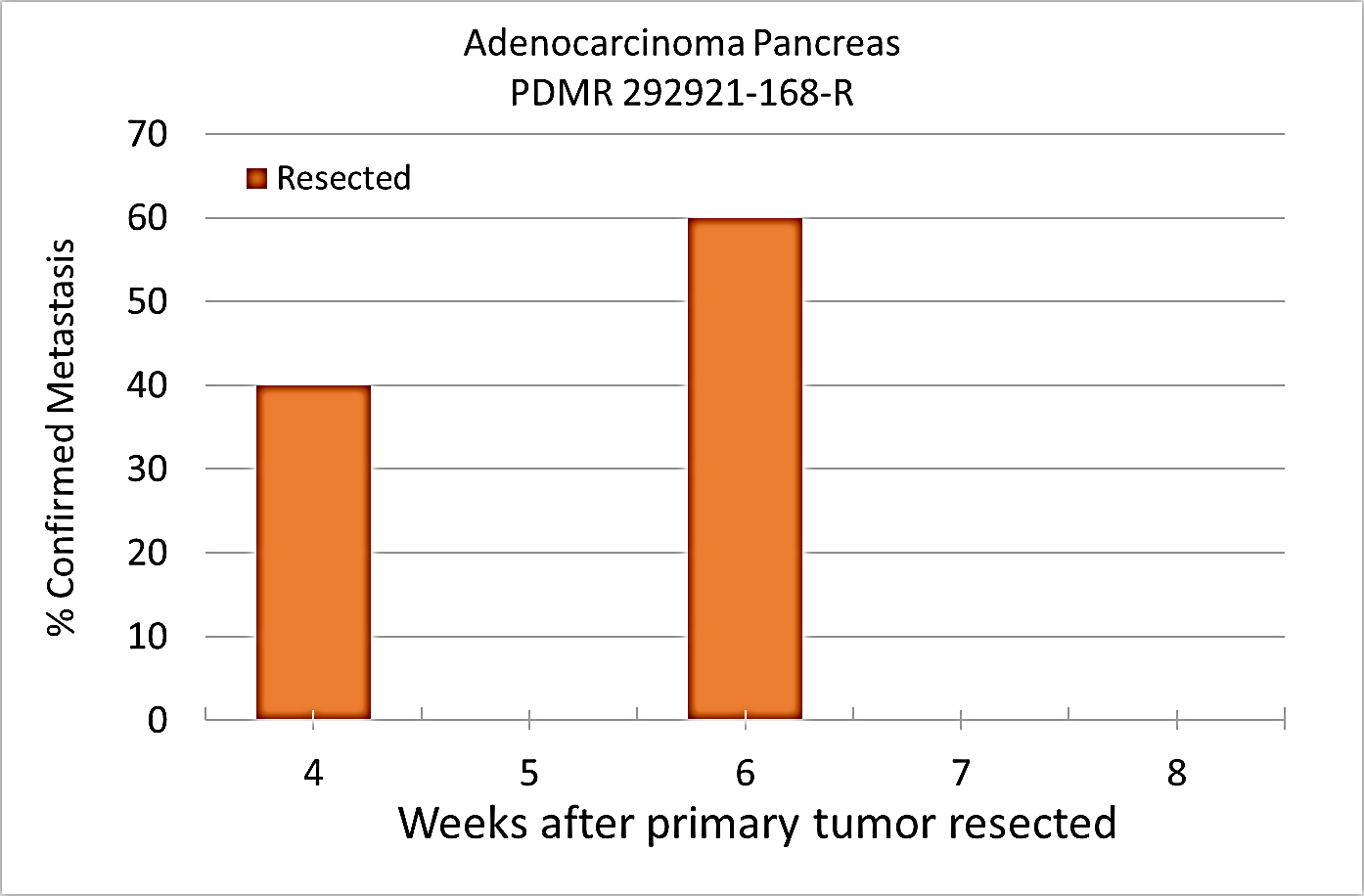
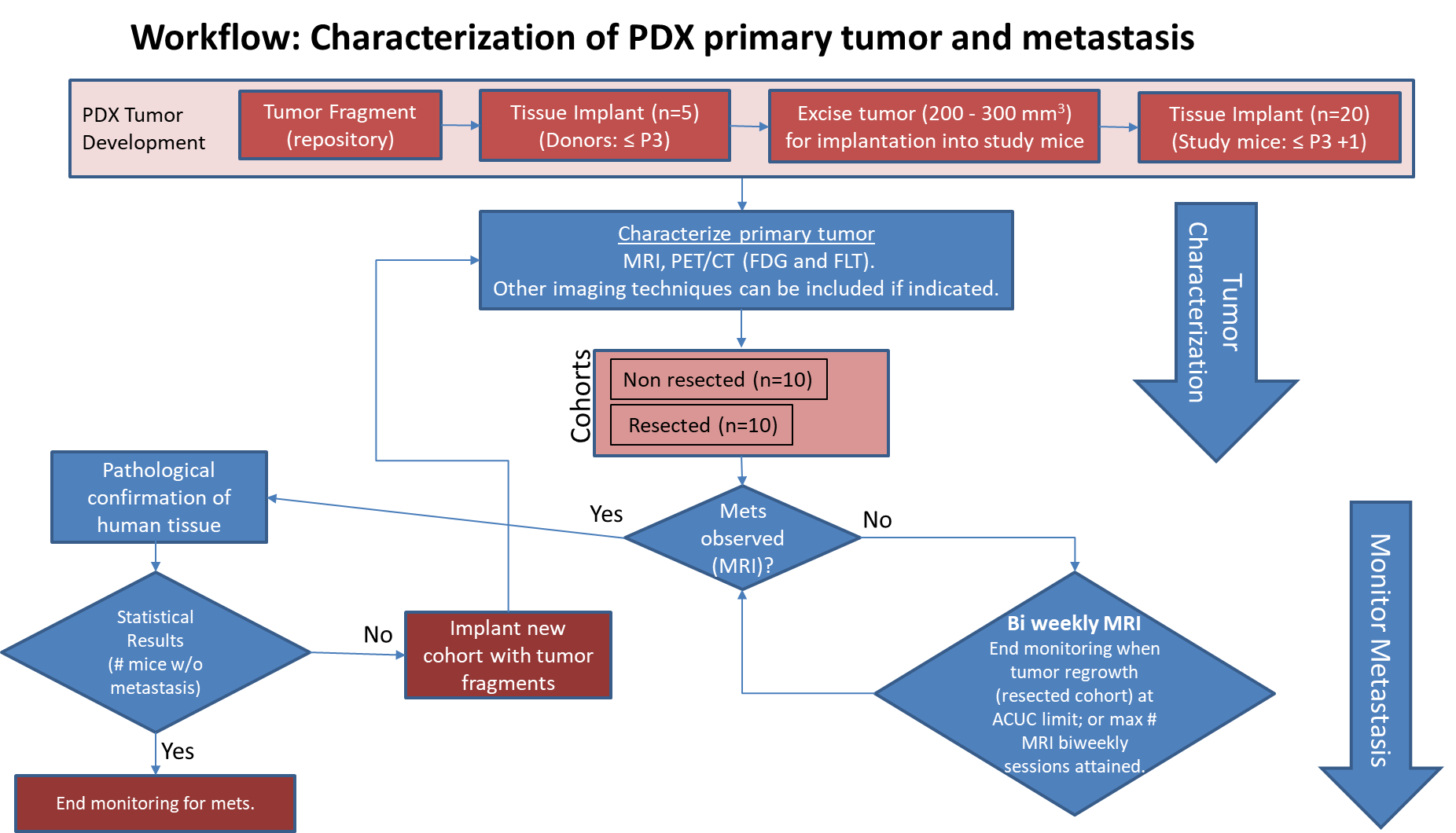
Pre-clinical animal models of spontaneous metastatic cancer are infrequent; the few that exist are resource intensive because determination of the presence of metastatic disease, metastatic burden, and response to therapy normally require multiple timed cohorts with animal sacrifice and extensive pathological examination. We identified and characterized a patient derived xenograft model with metastatic potential, adenocarcinoma pancreas xenograft 292921-168-R. In this study we performed a detailed imaging characterization (workflow below) of this model, which develops spontaneous lung metastases, details are provided in the attached standard operating procedures. Tumors in half of the mice were resected in the range 200-300 mm3 size; tumors in the other half were allowed to grow until it was necessary to euthanize them because of tumor size. The imaging characteristics of this model (PDMR-292921-168-R), which is available from the National Cancer Institute Patient-Derived Models Repository (https://pdmr.cancer.gov/), is highly favorable for preclinical research studies of metastatic disease when used in conjunction with non-contrast T2 weighted MRI. Table 1: Penetrance and location of pathological confirmed metastatic lesion(s). # animals in Group # animals that displayed metastasis in MRI and confirmed by Pathology Pathology confirmation of MRI (primary imaging site) Other confirmed Location (s) 10 (non-resected) 4 (5 mice were EU due to xenograft size prior to observation of metastases) Lung Lung 1409, 1410, 1418, 1423 10 (resected) 10 Lung Lung, Kidney, Nodes, Peritoneal Wall 1405, 1407, 1408, 1411, 1413,1415, 1416, 1419, 1420, 1422 Percent penetrance with respect to the average time-to-metastasis for non-resected (plot A: time from implant) and resected (plot B: time from tumor resection) cohorts. PET/CT Characterization of the primary tumor: Baseline PET (SOP available below) were performed when tumor reached an approximate 200 mm3. Average SUVmax values (n=5) were calculated; [18F]FDG: 1.3 ± 0.2 and [18F]FLT: 2.3 ± 0.6. Excellent metastatic model with at least 50% penetrance un-resected and 100% with planned early resection. Metastases are well observed on T2 MRI imaging allowing non-invasive evaluation in treatment trials.
Results: Adenocarcinoma pancreas (PDMR-292921-168-R)
Mouse ID: MRI with pathology confirmation of metastasis Conclusion
Data Access
Version 1: Updated 2020/03/23
| Title | Data Type | Format | Access Points | Subjects | License | Metadata | |||
|---|---|---|---|---|---|---|---|---|---|
| Images | MR, SR | DICOM | Download requires NBIA Data Retriever |
19 | 86 | 154 | 2,546 | CC BY 4.0 | View |
| Standard Operating Procedure 50101: MRI T2 Weighted Non-Contrast Protocol: Single Mouse Pulmonary Gated and Multi-Mouse Non-Gated | Protocol | CC BY 4.0 | — | ||||||
| Standard Operating Procedure 50102: Positron Emission Tomography imaging protocol | Protocol | CC BY 4.0 | — |
Additional Resources for this Dataset
The National Cancer Institute (NCI) has developed a national repository of Patient-Derived Models (PDMs) comprised of patient-derived xenografts (PDXs), in vitro patient-derived tumor cell cultures (PDCs) and cancer associated fibroblasts (CAFs) as well as patient-derived organoids (PDOrg). These models serve as a resource for public-private partnerships and for academic drug discovery efforts. These PDMs are clinically-annotated with molecular information and made available in the Patient-Derived Model Repository. Data related to the specific subjects in this Collection can be found at:
The NCI Cancer Research Data Commons (CRDC) provides access to additional data and a cloud-based data science infrastructure that connects data sets with analytics tools to allow users to share, integrate, analyze, and visualize cancer research data.
- Imaging Data Commons (IDC) (Imaging Data)
Citations & Data Usage Policy
Data Citation Required: Users must abide by the TCIA Data Usage Policy and Restrictions. Attribution must include the following citation, including the Digital Object Identifier:
Data Citation |
|
|
Tatum J., Kalen J., Ileva L.V., Riffle L.A., Saito K., Patel N., Jacobs P., Sanders C., James A., Difilippantonio S., Thang L., Hollingshead M.G., Phillips J., Evrard Y., Clunie D., Liu Y., Suloway C., Smith K.E., Wagner U., Freymann J.B., Kirby J., Doroshow, J. (2020). Imaging characterization of a metastatic patient derived model of adenocarcinoma pancreas: (PDMR-292921-168-R) [Dataset]. The Cancer Imaging Archive. https://doi.org/10.7937/TCIA.2020.PCAK-8Z10 |
Detailed Description
In addition to images, this collection includes Raw Data Storage SOP Class instances with MR Modality, generated by a Philips MR scanner; this data is not useful to anyone without the proprietary software to interpret it.
Acknowledgements
We would like to acknowledge the individuals and institutions that have provided data for this collection:
Frederick National Laboratory for Cancer Research – Special Thanks to Joseph D. Kalen, PhD, Lilia V. Ileva, MS, Lisa A Riffle, Nimit Patel, Keita Saito, PhD, Yvonne Evrard, PhD, Elijah Edmondson, DVM, PhD, Jessica Phillips, Simone Difilippantonio, PhD, Chelsea Sanders, Amy James, Lia Thang, Ulrike Wagner, Yanling Liu, PhD, John B. Freymann, and Justin Kirby.
Division of Cancer Therapeutics and Diagnosis/National Cancer Institute - James L. Tatum, MD, Paula M Jacobs, PhD, Melinda G. Hollingshead, DVM, and James H. Doroshow, MD
PixelMed Publishing – Special Thanks to David A. Clunie, MD
University of Arkansas for Medical Sciences – Special Thanks to Kirk E. Smith
This project has been funded in whole or in part with Federal funds from the National Cancer Institute, National Institutes of Health, under Contract Number HHSN261201500003I. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government.
Related Publications
Publications by the Dataset Authors
The authors recommended the following as the best source of additional information about this dataset:
No other publications were recommended by dataset authors.
Research Community Publications
TCIA maintains a list of publications that leveraged this dataset. If you have a manuscript you’d like to add please contact TCIA’s Helpdesk.