COVID-19-NY-SBU | Stony Brook University COVID-19 Positive Cases
DOI: 10.7937/TCIA.BBAG-2923 | Data Citation Required | Image Collection
| Location | Species | Subjects | Data Types | Cancer Types | Size | Status | Updated | |
|---|---|---|---|---|---|---|---|---|
| Lung | Human | 1,384 | MR, OT, SR, DX, NM, CR, CT, PT | COVID-19 (non-cancer) | Clinical, Image Analyses | Public, Complete | 2021/08/11 |
Summary
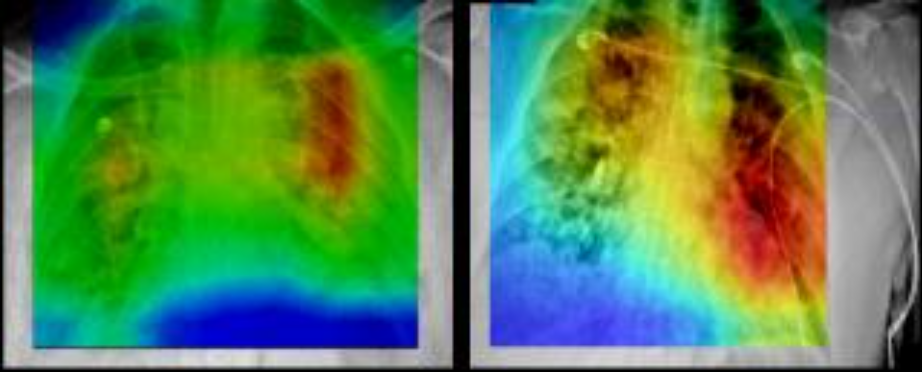
This collection of cases was acquired at Stony Brook University from patients who tested positive for COVID-19. The collection includes images from different modalities and organ sites (chest radiographs, chest CTs, brain MRIs, etc.). Radiology imaging data is extremely important in COVID-19 from both a diagnostic and a monitoring perspective, given the crucial nature of COVID-19 pulmonary disease and its rapid phenotypic changes. The datasets are available for building AI systems for diagnostic and prognostic modeling. This collection also includes associated clinical data for each patient. The clinical data consists of diagnoses, procedures, lab tests, covid19 specific data values (e.g., intubation status, symptoms at admission) and a set of derived data elements, which were used in analyses of this data. The clinical data is stored as a set of csv files which comply with OMOP Common Data Model data elements. The images on the right show automated identification of regions of prognostic importance on baseline chest radiographs. The regions of highest prognostic importance (as determined by the AI algorithm) are observed primarily in lower lung regions, consistent with clinical findings on the corresponding CXRs.
Data Access
Version 1: Updated 2021/08/11
| Title | Data Type | Format | Access Points | Subjects | License | |||
|---|---|---|---|---|---|---|---|---|
| Images | MR, OT, SR, DX, NM, CR, CT, PT | DICOM | Download requires NBIA Data Retriever |
1,384 | 7,361 | 17,950 | 562,376 | CC BY 4.0 |
| Clinical data | CSV | CC BY 4.0 | ||||||
| Clinical data template | CSV | CC BY 4.0 |
Additional Resources for this Dataset
The NCI Cancer Research Data Commons (CRDC) provides access to additional data and a cloud-based data science infrastructure that connects data sets with analytics tools to allow users to share, integrate, analyze, and visualize cancer research data.
Citations & Data Usage Policy
Data Citation Required: Users must abide by the TCIA Data Usage Policy and Restrictions. Attribution must include the following citation, including the Digital Object Identifier:
Data Citation |
|
|
Saltz, J., Saltz, M., Prasanna, P., Moffitt, R., Hajagos, J., Bremer, E., Balsamo, J., & Kurc, T. (2021). Stony Brook University COVID-19 Positive Cases [Data set]. The Cancer Imaging Archive. https://doi.org/10.7937/TCIA.BBAG-2923 |
Detailed Description
For a set of Covid+ patients (PCR positive), images were extracted from the Radiology PACS at Stony Brook Medicine and de-identified using POSDA. Images were matched with clinical data from the local Covid Data Commons. The Covid Data Commons is based on data captured from the electronic health records (EHR) at Stony Brook Medicine and manual review of clinical charts.
The main data file is named ‘deidentified_overlap_tcia.csv.cleaned.csv’. The file contains one row per patient whose images have been extracted. For each patient one encounter is selected using an algorithm (see “Encounter/visit selection steps” below for more detail). The algorithm is designed to select the Covid+ encounter where the patient had their most severe encounter. Images should be interpreted and aligned with the date-shifted field visit_start_datetime to correlate severity with the imaging data.
Clinical Data key
A description of fields in the de-identified files are provided in the file named ‘deidentified_overlap_tcia.csv.cleaned.csv.template.csv’. The column in the description file is_chart_abstracted indicates whether the column is derived from the manual chart review. Some field names are descriptive and so no additional information is provided. For laboratory and vital measurements the first value for the patient is selected.
Values of NA indicate that the value is missing, TRUE is a boolean True, FALSE is a boolean False. Original encoding from the source data of {Yes, No} are preserved in the final file. Some numeric measurement fields are constructed as: 2075-0_Chloride [Moles/volume] in Serum or Plasma where 2075-0 is the LOINC code and Chloride [Moles/volume] is the description associated with the LOINC code. LOINC codes and descriptions can be found on the LOINC website, for example, 2075-0.
Encounter/visit selection steps
The first steps of the algorithm is to find Covid+ patients and their potential encounters associated with infection:
- Apply date cut-off of February 1, 2020 for either the start or the end of an encounter.
- Remove future visits and remove any non-discharged (active) encounters.
- Identify the patient encounters where there are Covid+ PCR tests.
- Select visits which occur up to 7 days after the Covid+ PCR test.
- Identify Covid+ patients with encounters who have the ICD-10 code (U07.1) for Covid-19 virus identified.
In the second part of the algorithm we filter the encounters down to a single encounter – the most severe encounter:
- If a patient has only one encounter select this encounter.
- If a patient has multiple encounters, first select the inpatient encounters.
- If the patient has remaining encounters, select the hospital observation encounters.
- If the patient has remaining encounters, select the emergency department encounters.
- If the discharge disposition is death or hospice for an encounter, select that encounter and drop the others for that patient.
- If there is an encounter where the patient required invasive ventilation or ECMO, select that encounter.
- Pick the encounter with the longest length of stay.
- If there are still multiple encounters remaining for a patient, select the most recent one.
Acknowledgements
Data collection was enabled by the Renaissance School of Medicine at Stony Brook University’s “COVID-19 Data Commons and Analytic Environment”, a data quality initiative instituted by the Office of the Dean, and supported by the Department of Biomedical Informatics.
Related Publications
Research Community Publications
No research community publications were found.
